Histone modifications are chemical changes to proteins that influence gene activity without altering DNA. These modifications are vital for developing cell lines used in cultivated meat production, helping cells grow, maintain their identity, and differentiate into muscle tissue. The article explores how specific histone marks like H3K4me3 (gene activation), H3K27ac (enhancer activity), and H3K27me3 (gene repression) regulate cell behaviour.
Key points covered:
- H3K4me3 supports active genes and quick differentiation.
- H3K27ac controls enhancers for gene expression during growth phases.
- H3K27me3 ensures unwanted gene programmes stay inactive.
- Chromatin states, shaped by these marks, vary across species and cell types, affecting production quality.
The article also highlights recent research, including how positional gene expression in porcine cells impacts meat quality and how targeted epigenetic editing can improve cell line performance. Future directions include refining epigenetic tools and studying chromatin states to optimise production efficiency and scale.
Histone Modifications Explained | Acetylation, Methylation & Gene Regulation
sbb-itb-ffee270
Types of Histone Modifications and Their Functions
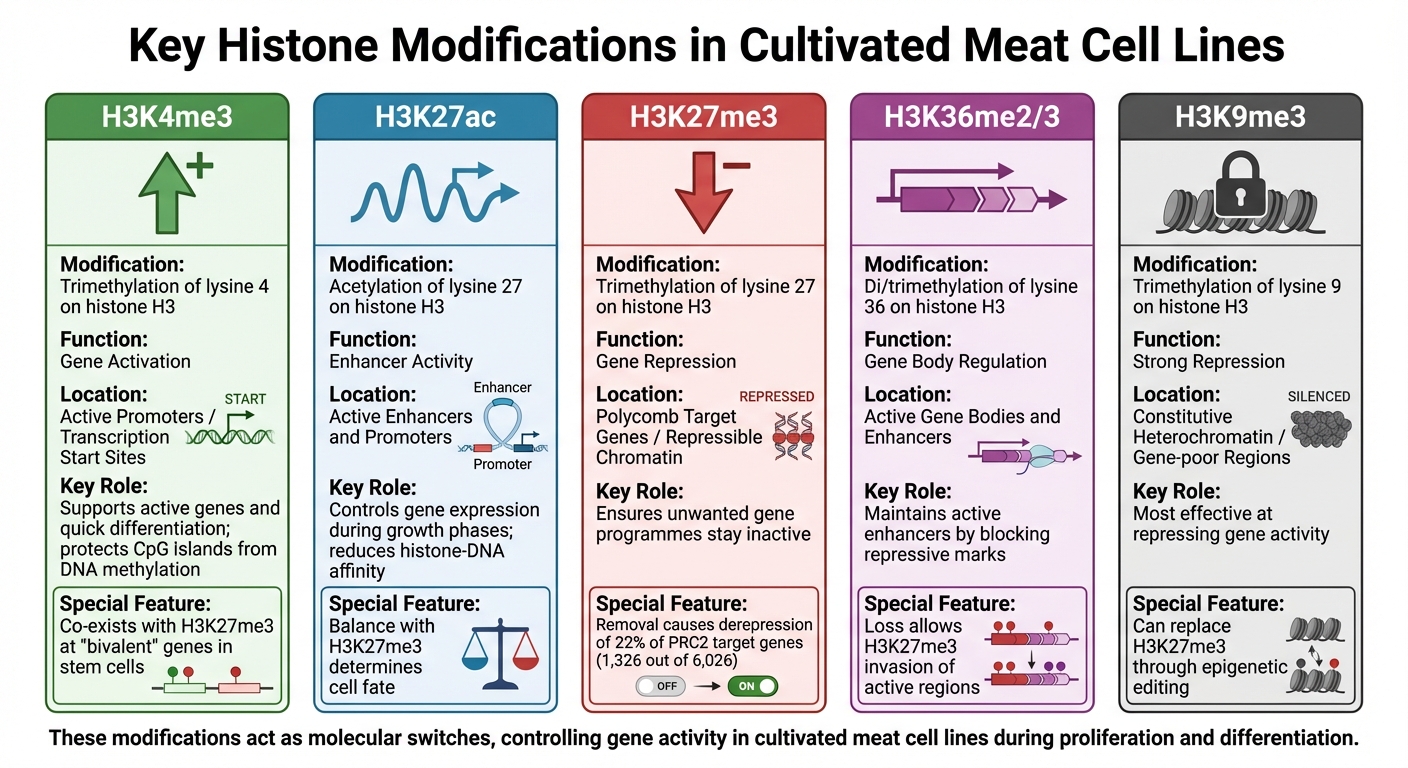
Key Histone Modifications in Cultivated Meat Cell Lines: Functions and Genomic Contexts
Histone modifications play a crucial role in regulating gene activity, acting like molecular switches to control whether genes are turned on or off in cultivated meat cell lines. These chemical tags - mainly methylation and acetylation - attach to specific residues on histones, creating distinct genomic patterns. Each modification has a specific function, and by understanding these roles, researchers can better predict and influence cell behaviour during production. This knowledge is essential for optimising processes in cultivated meat bioprocessing.
Here’s a breakdown of the main histone modifications that influence gene regulation in cultivated meat cell lines.
H3K4me3 and Gene Activation
H3K4me3 (trimethylation of lysine 4 on histone H3) is associated with active gene promoters and facilitates transcription at gene start sites, particularly for genes involved in cell growth and metabolism. This modification also protects CpG island promoters from new DNA methylation, ensuring essential genes stay accessible for transcription [4].
In primary or immortalised cell lines used for cultivated meat, H3K4me3 often co-exists with repressive marks like H3K27me3 at "bivalent" genes. These genes remain poised for activation, enabling rapid differentiation into muscle tissue when needed [4].
Interestingly, H3K4me3 interacts with other modifications. For instance, the deposition of H3K36me3 can inhibit H3K4 methyltransferases, reducing H3K4me3 levels at promoters and altering gene expression patterns [4].
H3K27ac and Enhancer Activity
H3K27ac (acetylation of lysine 27 on histone H3) is a marker of active enhancers and promoters. By reducing the affinity between histones and DNA, H3K27ac creates an environment that promotes transcription [5]. In cultivated meat cell lines, changes in H3K27ac levels during different growth phases determine which genes are expressed as cells move from proliferation to differentiation.
The balance between H3K27ac and repressive modifications like H3K27me3 is key to determining cell fate. For example, the loss of H3K36me2, which supports enhancer activity, can allow H3K27me3 to invade previously active regions, reducing H3K27ac levels and silencing target genes [5].
H3K27me3 and Gene Repression
H3K27me3 (trimethylation of lysine 27 on histone H3) is a repressive mark that promotes closed chromatin structures, effectively silencing genes. This modification, catalysed by the Polycomb Repressive Complex 2 (PRC2), is critical for maintaining the repression of thousands of developmental genes [4].
In cultivated meat cell lines, H3K27me3 ensures that unwanted gene programmes remain inactive during specific growth phases, preserving the cells' intended identity.
"H3K27me3, together with H2AK119ub1, is essential for maintaining the transcriptional repression of several thousand Polycomb target genes." - Nature Communications [4]
Research has shown that removing H3K27me3 in mouse embryonic stem cells results in the derepression of about 22% (1,326 out of 6,026) of PRC2 target genes [4]. For cultivated meat, controlling this modification can help suppress alternative cell fates, such as fat or connective tissue formation, while focusing on muscle development.
| Histone Modification | Regulatory Function | Genomic Context |
|---|---|---|
| H3K4me3 | Gene Activation | Active Promoters / Transcription Start Sites |
| H3K27ac | Enhancer Activity | Active Enhancers and Promoters |
| H3K27me3 | Gene Repression | Polycomb Target Genes / Repressible Chromatin |
| H3K36me2/3 | Gene Body Regulation | Active Gene Bodies and Enhancers |
| H3K9me3 | Strong Repression | Constitutive Heterochromatin / Gene-poor Regions |
Chromatin States in Cultivated Meat Cell Lines
Histone modifications don’t act alone - they combine to form chromatin states, which are unique genomic environments that control gene accessibility. These states play a crucial role in shaping the behaviour of cultivated meat cell lines during both expansion and differentiation, making them key to optimising bioprocessing.
Identifying Chromatin States Through Histone Marks
Researchers map chromatin states by studying combinations of histone marks like H3K4me3, H3K27ac, and H3K27me3. For instance, in porcine foetal fibroblasts (PFF) and trophectoderm (PTr2) cells, 10 distinct chromatin states have been identified, including active transcription start sites, bivalent promoters, and putative enhancers [6]. These states help predict gene activity.
Enhancer states, marked predominantly by H3K27ac in intergenic and intronic regions, are often co-enriched with the chromatin remodelling protein BRG1 [6].
A particularly notable feature is the presence of broad H3K4me3 domains, which span regions of 4 kb or more. These domains represent just 1.7% to 1.8% of all predicted transcription start sites in porcine cell lines but are critical for marking developmental and tissue-specific genes [6]. Interestingly, in porcine foetal fibroblasts, 52% of genes marked by these broad domains are tissue-specific, compared to only 25% in PTr2 cells [6].
"These findings enhance our understanding of the epigenetic landscape present in early swine development and provide insight into how variabilities in chromatin state are linked to cell identity." - BMC Epigenetics & Chromatin [6]
These chromatin state profiles not only differ within a single species but also vary across the various animal cell lines used in cultivated meat production.
Chromatin Differences Across Animal Cell Lines
Chromatin state patterns change significantly depending on the species and cell type used in cultivated meat production. For example, in chicken cell lines, H3K4me3 accounts for 30% to 55% of its genomic presence at gene promoters [7]. However, in chicken primordial germ cells (PGCs), H3K4me3 levels drop substantially compared to pluripotent cells. This reduction supports the transition of bivalent states into repressive states during germline specification [7].
Porcine trophectoderm (PTr2) cells show higher H3K27ac levels in promoter regions (57.36%) compared to foetal fibroblasts (41.58%), while H3K27me3 enrichment is lower in PTr2 cells (7.77%) than in PFF cells (22%) [6]. These variations reflect the distinct epigenetic needs of each developmental stage and influence how these cells respond to culture conditions.
In bovine satellite cells, differentiation into a "reserve cell" fate (Pax7+/Ki-67-) is driven by quiescent chromatin states regulated by NOTCH and MAPK/ERK signalling. This process, however, reduces protein yield [3]. Such variability underscores how chromatin states directly impact production efficiency. Gaining a deeper understanding of these differences is crucial for fine-tuning cell line performance in cultivated meat production.
Using Histone Modifications to Improve Cell Lines
Building on what we know about chromatin states, let’s delve into how targeted histone modifications can directly improve the performance of cultivated meat cell lines.
Boosting Proliferation and Adaptation to Suspension Growth
Adjusting histone marks can significantly increase cell proliferation and help cells transition from adherent to suspension growth. This shift is crucial for bioreactor systems for cultivated meat. For example, reducing H3K36 methylation makes fibroblasts less responsive to TGFβ, resulting in a more flexible cell state [1].
In December 2022, researchers at Believer Meats achieved a breakthrough with chicken fibroblasts (HUN-CF-2 and HUN-CF-4). They demonstrated spontaneous immortalisation in serum-free suspension cultures, reaching 100 million cells per ml (10⁸ cells/ml) and achieving yields of 36% w/v. The team, led by Yaakov Nahmias, used lecithin - a food-safe small molecule - to activate the PPARγ pathway and promote fat formation without relying on genetic modification. Their cultivated chicken prototype earned a sensory rating of 4.5 out of 5.0 [2].
"Immortalisation without genetic modification and high-yield manufacturing are critical for the market realisation of cultured meat." - Yaakov Nahmias, Chief Scientific Officer, Believer Meats [2]
These findings highlight the potential of precise epigenetic tools to further refine cell line development.
Precision in Epigenetic Editing
To complement these cellular changes, precise epigenetic editing methods allow for targeted manipulation of histone marks. A 2025 study on mouse embryonic stem cells showed that a chimeric recruiter (S12N) fused with catalytic domains from SUV39H2 or SETD2 could replace H3K27me3 with H3K9me3 or H3K36me3 at thousands of genes. Among these, H3K9me3 proved to be more effective at repressing gene activity [8].
However, the success of these modifications depends heavily on the existing chromatin environment. For instance, residual H3K4me3 at gene promoters can block DNA methylation machinery, making it harder to achieve the desired gene silencing [8]. This suggests that optimising cell performance often requires adjusting multiple histone marks at the same time rather than focusing on a single modification.
Conclusion and Future Directions
Key Takeaways
Histone modifications play a critical role as molecular switches, controlling gene activity in cultivated meat cell lines. Specifically, H3K36me2 and H3K36me3 help maintain active enhancers by blocking repressive marks like H3K27me2/3 from infiltrating gene bodies [9][10]. When H3K36 methylation is lost, chromatin structure is disrupted, allowing repressive marks such as H3K9me3 to invade active regions [9].
"H3K36 methylation [is] a pivotal regulator of chromatin state and genomic structure." - Nature Communications [9]
The interaction between histone marks is essential for improving cell line performance. Research suggests that targeting multiple histone modifications together often achieves better outcomes than focusing on just one [4].
With these findings in mind, future studies must leverage precision epigenetic tools to ensure ongoing improvements in cultivated meat cell line performance.
Future Research Opportunities
Advancing cell line performance calls for innovative approaches, such as single-nucleus RNA sequencing, to map the epigenetic landscape within different cell subpopulations. This is particularly crucial for identifying and understanding quiescent "reserve cells" that resist differentiation. These cells, which express markers like PAX7 and NOTCH2 instead of committing to myogenic fusion, present a significant challenge in cultivated meat production [3].
Another promising avenue involves the development of chimeric epigenetic complexes for precise, non-genetic control. For example, in 2025, researchers demonstrated that combining the N-terminal of SUZ12 with catalytic domains from SUV39H2 or SETD2 could effectively replace H3K27me3 with H3K9me3 or H3K36me3 at numerous genes [4]. Additionally, monitoring H3K36me2 at enhancers could act as a quality control marker to ensure cell line stability [9].
Future efforts should focus on maintaining H3K36 methylation across cell generations. This would help prevent epigenetic drift, enabling researchers and companies like Cellbase to profile these marks for consistent performance in bioreactor systems for large-scale production. By addressing these challenges, the cultivated meat industry can move closer to achieving reliable and scalable production.
FAQs
How do histone marks affect muscle differentiation in cultivated meat cell lines?
Histone marks are key players in muscle differentiation, particularly for cultivated meat cell lines. For example, the reduction of H3K27me3 during differentiation triggers myogenic transcription programmes, allowing the activation of genes necessary for muscle development. Fine-tuning histone modifications like H3K27me3 supports the transition of cell lines from proliferation to forming muscle tissue with specific characteristics. These epigenetic adjustments are essential for advancing the production of cultivated meat.
Which histone modifications best predict stable, high-yield cell growth in bioreactors?
H3K36 methylation stands out as a dependable marker for stable, high-yield cell growth in bioreactors. This modification plays a key role in preserving cell identity and managing lineage programmes - both of which are essential for ensuring consistent cell proliferation, particularly in the production of cultivated meat.
Can epigenetic editing improve cell lines without changing their DNA sequence?
Epigenetic editing offers a way to improve cell lines without changing their DNA sequence. By adjusting histone marks and chromatin structure, it controls gene expression. Research on histone modifications highlights how these alterations can affect cell identity and function. This approach holds particular promise for refining cultivated meat cell lines.











