High-throughput CRISPR screening is transforming the cultivated meat sector by enabling precise genetic modifications to improve cell line performance. Here's what you need to know:
- Key Challenge: Cultivated meat production requires cell lines that grow efficiently, resist ageing, and differentiate into muscle and fat tissue.
- CRISPR's Role: By targeting thousands of genes simultaneously, these platforms identify genetic edits that enhance growth, delay senescence, and support differentiation.
- Notable Findings: Studies have shown that knocking out genes like TP53 and PTEN in bovine mesenchymal stem cells can increase proliferation up to 1,000-fold in 30 days and extend their lifespan from 100 to 200 days.
- Applications: CRISPR tools such as knockout screens, CRISPRi, and CRISPRa are being used to optimise cell growth, regulate gene expression, and balance proliferation with differentiation.
- Industry Tools: Advanced techniques like RMCE, RNA-seq, and single-cell platforms integrate CRISPR results with multi-omics data, ensuring precise and scalable improvements.
For bioprocess engineers and R&D professionals, these innovations address critical bottlenecks in scaling cultivated meat processes while maintaining cell quality and functionality. The integration of CRISPR with automated systems and tailored resources like Cellbase further accelerates industrial viability.
CRISPR-Cas9 Fundamentals for Genome-Wide Knockout Screens
How CRISPR-Cas9 Works in Large-Scale Gene Editing
The CRISPR-Cas9 system relies on a Cas9 nuclease paired with a single-guide RNA (sgRNA) to target specific DNA sequences. Once the sgRNA directs Cas9 to the desired genomic location, the enzyme creates a double-strand break in the DNA. This break is predominantly repaired via non-homologous end joining (NHEJ), an error-prone process that often introduces small insertions or deletions (indels). These indels can cause frameshift mutations, effectively disrupting the function of the targeted gene [1]. This precise mechanism is the foundation for conducting genome-wide knockout screens, which are instrumental in identifying critical regulators of cellular behaviour.
For large-scale screening, researchers use a diverse library of sgRNAs, typically delivered into a mixed cell population through lentiviral transduction. To ensure that each cell receives only one genetic alteration, a low multiplicity of infection (MOI of around 0.3) is maintained [1]. Over time, cells with advantageous mutations tend to proliferate more successfully than others, a phenomenon observed across a variety of cell types and experimental conditions.
Alternative delivery methods, such as recombinase-mediated cassette exchange (RMCE), offer additional precision by targeting specific genomic "landing pads" to reduce variability in integration sites. For example, a study using CHO-K1 cells employed a virus-free RMCE method to screen 111,651 unique gRNAs across 21,585 genes. This approach identified genes essential for cell fitness over 16- and 37-day periods [7].
Benefits of Genome-Wide Screening
Genome-wide knockout screens leverage the accuracy of CRISPR-Cas9 to systematically investigate thousands of genes. This enables researchers to uncover genes that influence cell survival, growth, and responses to stress. Beyond genetic factors, optimizing surface functionalization is critical for improving cell attachment and growth in these systems. Such unbiased exploration is especially relevant to cultivated meat production, where mesenchymal stem cells (which make up about 25% of cell sources) often face challenges like limited proliferation and early senescence [1].
sbb-itb-ffee270
Pooled CRISPR Library Screening Methods
Building Pooled CRISPR Libraries
Pooled CRISPR libraries start with a carefully selected collection of single-guide RNAs (sgRNAs). In the context of cultivated meat research, targeted libraries are often designed to focus on specific gene families, such as transcription factors or regulators of cell proliferation. This approach helps balance cost with scalability while keeping the focus on traits relevant to the desired phenotype [1].
The process begins with synthesising oligonucleotides as a pool, amplifying them via PCR, and cloning them into a delivery vector. For instance, a bovine-specific library constructed in early 2025 included 3,000 sgRNAs targeting 603 genes to identify factors influencing stem cell expansion [1]. On a larger scale, genome-wide screens can reach much higher complexity. One example is a Chinese Hamster Ovary (CHO) cell screen, which used 111,651 unique gRNAs to target 21,585 genes [7].
Lentiviral transduction is commonly used to deliver these libraries at a low multiplicity of infection (around 0.3), ensuring each cell undergoes only a single genetic modification [1]. Alternatively, virus-free methods like recombinase-mediated cassette exchange (RMCE) integrate the gRNA library into predetermined genomic "landing pads" within a master cell line. This technique achieves 99.9% gRNA coverage with minimal skewing [7].
To maintain statistical reliability, researchers ensure high coverage - typically 500 to 600 cells per sgRNA [1] [7]. Some platforms use inducible Cas9 (iCas9) systems, enabling precise control over when gene editing occurs. For example, editing can be triggered after cells reach a specific state, such as high density or the onset of senescence. This temporal control is particularly useful for studying non-proliferative phases, which are crucial for overcoming senescence barriers by choosing between primary vs immortalised cell lines for scaling cultivated meat production [4].
Once the library is constructed, researchers move on to targeted screening assays to evaluate gene function.
Screening Approaches for Cultivated Meat Cell Lines
After building the library, researchers assess cell performance using competition assays and functional sorting techniques. A widely used method is the competition-based proliferation assay, which identifies genetic changes that confer growth or senescence resistance - key traits for optimising cell lines for cultivated meat.
Short-term screens (lasting about 30 days) identify genes that immediately influence the cell cycle, while long-term screens (up to 200 days) focus on genes that help cells overcome replicative senescence. This is a critical challenge in scaling cultivated meat production [1]. For more complex traits, such as enhanced protein secretion or expression of specific markers, fluorescence-activated cell sorting (FACS) is employed. One example is the "cold capture secretion assay", which isolates productive cell populations by capturing secreted proteins on the cell surface before sorting [7] [5].
Validation is a crucial step in confirming screening results. The Cellular Fitness (CelFi) assay, for instance, tracks the ratio of out-of-frame to in-frame mutations over time. If cells with out-of-frame mutations disappear from the population, it suggests the targeted gene is essential for cellular fitness [2].
In June 2025, researchers led by Shijie Ding at Nanjing Agricultural University used CRISPR/Cas9 to create CDKN2A–/– porcine satellite cell lines. These modified cells maintained stable proliferation for at least 15 passages in serum-free conditions while retaining stemness markers. When seeded onto a plant-based 3D edible scaffold, they formed meat-like constructs with improved texture, including enhanced chewiness and gumminess [8].
"These findings demonstrate the utility of CRISPR screening for optimising bovine stem cell traits and offer a path toward more scalable cultured meat production in the future." – Communications Biology [1]
Pooled CRISPR-Genetic Screens in Mammalian Cells | Protocol Preview
CRISPRi and CRISPRa for Reversible Gene Regulation Screens
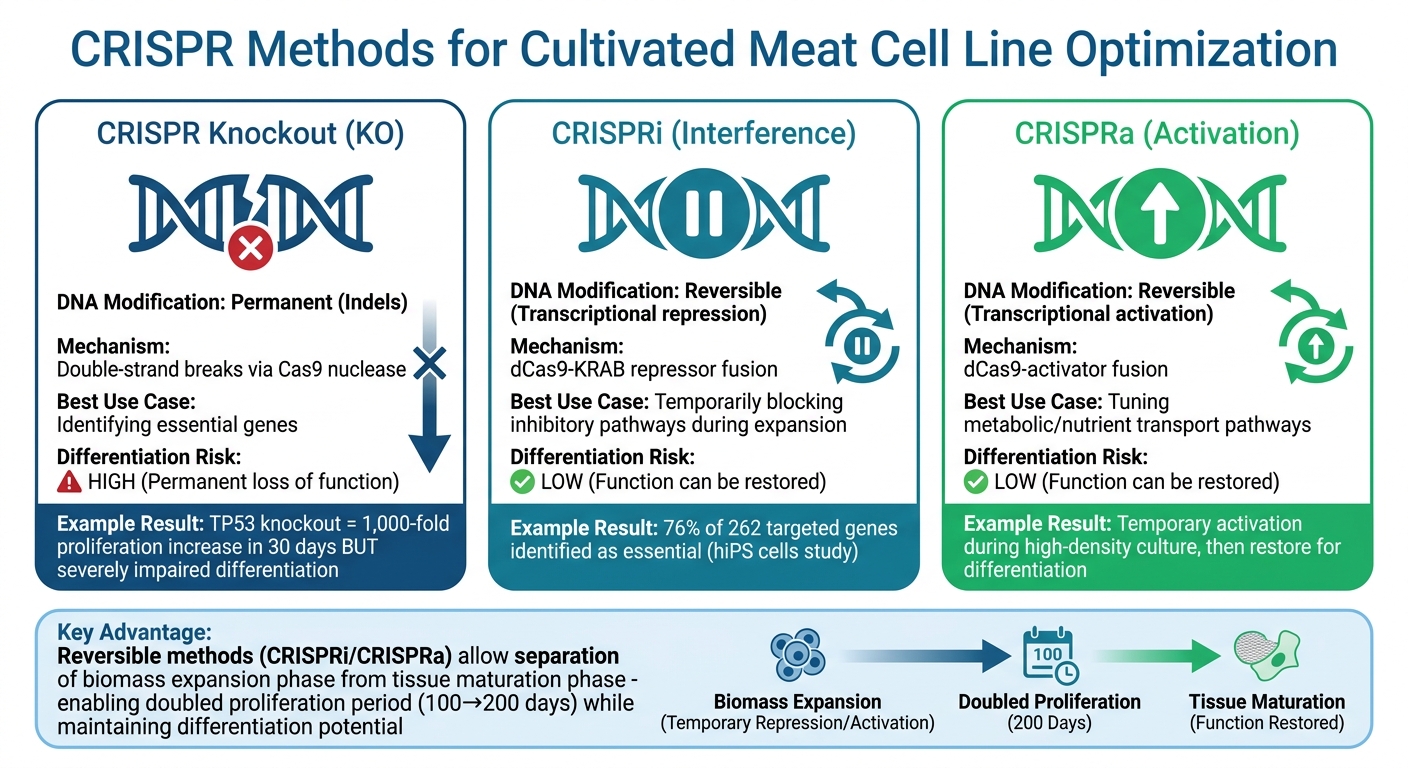
CRISPR Gene Editing Methods for Cultivated Meat: Knockout vs CRISPRi/CRISPRa Comparison
Using CRISPRi and CRISPRa in Functional Genomics
When it comes to improving cultivated meat production, CRISPR interference (CRISPRi) and CRISPR activation (CRISPRa) provide powerful tools. These techniques use an inactive Cas9 protein paired with repressors or activators, allowing researchers to adjust gene expression temporarily without making permanent changes to the DNA [10].
This reversibility is especially important for tackling a major challenge: genes that promote rapid cell growth often interfere with the later stages of differentiation into muscle or fat tissue. For instance, permanently knocking out the TP53 gene in bovine mesenchymal stem cells can increase proliferation over 1,000-fold in just 30 days but severely impairs their ability to differentiate [1]. CRISPRi offers a more flexible solution by temporarily blocking pathways that inhibit differentiation during biomass expansion in bioreactors for cultivated meat. Once the cells are ready for tissue maturation, normal gene function can be restored.
In October 2025, researchers like Gabriele Casagrande Raffi and Roderick L. Beijersbergen from the Netherlands Cancer Institute developed an inducible CRISPR system. This approach delays gene editing until cells reach specific states - such as high density or a non-proliferative phase - helping to preserve cell viability [4].
CRISPRi also stands out for its precision compared to traditional RNA interference (RNAi). RNAi often leads to inconsistent results and off-target effects, while CRISPRi delivers more reliable and specific gene repression [2]. Another advantage is that CRISPRi avoids triggering p53-related toxicity, which is often caused by DNA damage responses. In a 2025 study led by Liqin Wang at Sun Yat-sen University Cancer Centre, researchers used a doxycycline-inducible KRAB–dCas9 system to screen 262 genes in human induced pluripotent stem cells (hiPS cells). They found that 76% of targeted translation-related genes (200 out of 262) were essential for growth, demonstrating the system's effectiveness [10].
This ability to fine-tune gene expression makes CRISPRi and CRISPRa valuable tools for balancing cell proliferation and differentiation in functional genomics research.
Adapting Reversible Screens for Cultivated Meat Applications
Reversible gene regulation offers solutions to key challenges in cultivated meat production. For example, CRISPRa can temporarily activate genes involved in nutrient transport or metabolic pathways during high-density culture. Once the cells reach the desired density, the system can return gene expression to normal levels, supporting proper differentiation into muscle or fat tissue.
Inducible systems also make it possible to separate the biomass expansion phase from tissue maturation. CRISPRi can suppress senescence-associated genes during the scale-up process, effectively doubling the proliferation period of bovine cells from around 100 days to over 200 days [1]. After achieving sufficient biomass, researchers can restore normal gene expression to enable differentiation. This approach is particularly useful for mesenchymal stem cells, which tend to enter senescence early in culture [1].
"Targeted genetic editing of these dual processes could optimise MSC expansion efficiency whilst maintaining their essential multipotency and differentiation potential, ultimately advancing scalable cultured meat systems." – Communications Biology [1]
The table below highlights the differences between reversible and permanent gene regulation methods:
| Feature | CRISPR Knockout (KO) | CRISPRi / CRISPRa |
|---|---|---|
| DNA Modification | Permanent (Indels) | Reversible (Transcriptional) |
| Mechanism | Double-strand breaks | dCas9-effector fusion |
| Best Use Case | Identifying essential genes | Tuning metabolic/growth pathways |
| Differentiation Risk | High (Permanent loss of function) | Low (Function can be restored) |
This comparison illustrates how reversible gene regulation methods can be tailored to meet the specific challenges of developing cell lines for cultivated meat production.
Combining CRISPR Screens with Cell Panel and Genotyping Technologies
Linking CRISPR Screens with Multi-Omics Analysis
Integrating multi-omics and automated genotyping into CRISPR screens refines their utility, particularly in advancing cultivated meat cell line development.
Combining CRISPR screens with multi-omics, such as RNA sequencing, allows researchers to map the effects of specific gene knockouts on cellular pathways. This is especially relevant for cultivated meat, where understanding how cells balance proliferation and differentiation is critical.
For example, a pooled CRISPR knockout screen targeting 600 genes in bovine adipose-derived mesenchymal stem cells, paired with RNA-seq, uncovered that knocking out TP53 and PTEN delayed senescence. These cells maintained a youthful gene expression profile, with upregulated cell-cycle genes, leading to a 50% increase in doubling rates by day 50 post-transduction [1].
Single-cell platforms like CROP-seq take this further by simultaneously detecting both sgRNA and transcriptomic changes in individual cells [6]. This level of precision is invaluable for identifying genetic modifications that enhance muscle differentiation or protein synthesis - critical factors for achieving the desired texture and nutritional properties in cultivated meat.
Another promising approach involves cell panel screening, where CRISPR perturbations are tested across diverse cell lines from various donors, anatomical sites, and species. For instance, researchers validated the MyoCRISPR-KOLib library on human myoblast lines from seven donors. Using a split-toxin selection system, they identified 250 genes essential for myoblast fusion. Of these, 41 genes were confirmed through medical databases to play a role in skeletal muscle morphology [6]. This multi-line validation ensures that genetic targets remain robust across biological variations, a key consideration for scaling cultivated meat production.
These insights are paving the way for automated, scalable platforms that combine genetic screens with detailed genotyping for industrial applications.
Automation and Scalability in Integrated Platforms
Automation is essential for handling the vast datasets and samples generated by integrated CRISPR and genotyping platforms. RMCE systems, which enable virus-free, site-specific delivery of sgRNA libraries, are a major step forward. These platforms ensure each cell receives a single, consistent sgRNA copy, reducing variability. RMCE has already demonstrated high library coverage with minimal bias in Chinese hamster ovary (CHO) cells [5].
"An unbiased high-throughput genetic screening platform is essential for the development of next-generation CHO factories." - Chinese Hamster Ovary Research Team [5]
Scalability is further enhanced by validation tools like the Cellular Fitness (CelFi) assay. This assay uses targeted deep sequencing to monitor indel profiles over time, tracking the ratio of in-frame versus out-of-frame mutations. By correlating these mutations with growth advantages or disadvantages, researchers can efficiently verify genetic targets in cultivated meat cell lines [2].
| Technology | Integration Method | Primary Benefit for Cultivated Meat |
|---|---|---|
| RNA-seq / Multi-omics | Linking CRISPR hits to transcriptomic profiles | Understanding how genes regulate growth and differentiation [1][6] |
| Split-Toxin Systems | Linking cell fusion to viability selection | Quantitative selection of fusion-capable or defective cells [6] |
| RMCE Platforms | Site-specific integration of gRNA libraries | High-throughput, virus-free screening with consistent gene copy numbers [5] |
| CROP-seq | Single-cell CRISPR + RNA-seq | Simultaneous detection of sgRNA and transcriptomic changes [6] |
| CelFi Assay | Targeted deep sequencing of indels | Fast validation of genetic targets by tracking allele frequency shifts [2] |
These advanced platforms streamline the process from identifying genetic targets to validating their impact on cell fitness. This efficiency supports the development of cell lines robust enough for large-scale cultivated meat production.
Using CRISPR Screens to Improve Cell Line Growth and Proliferation
CRISPR screening methods have become a powerful tool for enhancing cell line performance, offering direct benefits for cultivated meat production.
Examples of CRISPR-Based Cell Line Improvements
CRISPR screens have successfully improved cell line performance in cultivated meat research. For example, a pooled knockout screen targeting 600 genes in bovine adipose-derived mesenchymal stem cells identified TP53 and PTEN as key inhibitors of growth. Knocking out TP53 significantly increased cell abundance within 30 days[1]. Additionally, the edited bovine mesenchymal stem cells showed a 12% higher doubling rate on average[1].
By targeting tumour suppressor genes, researchers extended the cells' proliferative lifespan from about 100 to over 200 days, effectively bypassing the Hayflick limit. This delay in senescence enables biomass expansion over industrially relevant timeframes[1].
In another example, researchers from Nanjing Agricultural University, led by Shijie Ding, Chunbao Li, and Guanghong Zhou, used CRISPR/Cas9 to develop CDKN2A−/− porcine satellite cell lines. These engineered cells maintained stable proliferation for at least 18 passages in a custom 19-component serum-free medium (A19). They were also successfully seeded onto edible scaffolds, creating meat-like constructs with improved chewiness and gumminess[8]. The cells maintained over 90% viability across multiple passages in serum-free conditions[8].
"The CRISPR-based CDKN2A knockout cells provide a renewable source of muscle progenitors, reducing dependence on repeated animal biopsies."
These examples highlight how CRISPR screens can identify genetic modifications that improve growth rates, delay cellular ageing, and enable serum-free culture - three essential aspects for scaling cultivated meat production.
Scaling Challenges for CRISPR-Optimised Cell Lines
While CRISPR-optimised cell lines show clear advantages, scaling them for industrial use presents challenges. Enhanced proliferation often comes at the cost of differentiation. For instance, TP53 knockouts in bovine mesenchymal stem cells have been associated with reduced expression of muscle-differentiation genes, which can hinder their ability to mature into edible tissue[1]. To address this, additional strategies, such as adding media supplements or activating specific transcription factors, may be needed to restore differentiation after expansion[1].
Another critical issue is maintaining genetic stability. Variations in gene copy numbers (aneuploidy) and off-target effects during CRISPR editing can lead to inconsistent results or false positives in screening studies[2]. Tools like the Cellular Fitness (CelFi) assay help mitigate these risks by monitoring the ratio of out-of-frame indels over time, ensuring that observed growth benefits are directly linked to the intended edits[2].
Economic and technical barriers also remain. Mesenchymal stem cells, which make up about 25% of cell sources in the cultivated meat industry, face challenges such as the high cost of growth factors, the need for optimised serum-free media, and the development of large-scale bioreactors (10,000–50,000 L capacities)[1][9][11]. Additionally, ensuring the desired texture when cells are seeded onto 3D scaffolds continues to be a complex task[11].
"The current state of cultivated meat faces significant challenges, including high costs, scalability issues, and the need for further technological advancements."
- Communications Biology [1]
Overcoming these challenges requires a comprehensive approach that combines genetic optimisation with advancements in media formulation, bioreactor technology, and differentiation protocols. While CRISPR screens provide critical genetic insights, translating these findings into scalable solutions will demand integrated systems and rigorous validation processes. These efforts are vital for moving cultivated meat production from the lab to commercial viability.
How Cellbase Supports CRISPR Research in Cultivated Meat
CRISPR screening has already shown its potential, but scaling it for industrial use demands access to specialised tools and resources. This is where Cellbase steps in. As the first B2B marketplace dedicated to cultivated meat, Cellbase connects researchers with the critical CRISPR screening materials and cell line development resources they need.
Accessing CRISPR Resources Through Cellbase
Cellbase simplifies the procurement process for CRISPR-related inputs by centralising access to essential materials like animal cell lines (bovine, porcine, avian, and seafood), growth media, bioreactors, and lab equipment. Supplier data is standardised into structured fields, making it easy for researchers to compare options directly without sifting through disparate formats or documents [13]. This efficiency eliminates the need for manual reviews, saving valuable time.
Unlike broad-spectrum pharmaceutical suppliers, Cellbase offers products specifically validated for food production. This includes regulatory notes and compatibility information, which are crucial for scaling cultivated meat research [12][15]. For example, growth media - responsible for over 95% of cultivated meat production costs - can be sourced transparently through the platform, ensuring cost-effective and reliable procurement [12]. Additionally, the platform replaces the traditional "request for quote" system with transparent unit pricing and one-click ordering, significantly cutting down on administrative delays [13].
In November 2025, Cellcraft Ltd partnered with Cellbase to launch its first products on the platform. Under the leadership of Cultigen Group founder David Bell, the partnership focused on easing the procurement of specialised biomarkers and cell culture media for researchers in the UK. This collaboration tackled supply chain fragmentation by connecting producers with specialised suppliers like Multus, Sallea, and Quest Meat [13][14].
"Every cultivated meat company we spoke to was wasting time on the same procurement headache. Finding suppliers for critical components meant googling through pages of pharma suppliers who didn't understand food applications."
- David Bell, Founder of Cultigen Group [15]
By centralising these resources, Cellbase not only accelerates procurement but also sets the stage for a more unified research environment, encouraging global collaboration. This approach directly supports the advancement of CRISPR-driven cultivated meat development.
Enabling Collaboration in Cultivated Meat Development
Cellbase goes beyond procurement by fostering international partnerships. Supporting 20 languages and all major currencies, it serves around 95% of the global cultivated meat market [13]. This global accessibility allows researchers, suppliers, and companies to collaborate across borders seamlessly.
The platform is designed to handle the demands of large-scale commercial projects, like those undertaken by Believer Meats and Aleph Farms. These ventures require infrastructure for 50,000-litre bioreactors and optimised production supply chains, which Cellbase is equipped to support [12][15]. By connecting stakeholders worldwide, Cellbase plays a vital role in advancing cultivated meat research and production.
Conclusion
High-throughput CRISPR screening has transitioned from a promising concept to a critical tool in advancing cultivated meat development. The impact of this technology on optimising cell lines is undeniable. For instance, recent breakthroughs have shown that genetic modifications can double the proliferation lifespan of bovine stem cells from 100 to 200 days, slash senescent cell populations from 60% to just 10%, and achieve a staggering 1,000-fold increase in cell abundance within a single month [1]. These advancements mark a clear shift from experimental research to practical industrial applications.
Compact platforms and targeted libraries are addressing some of the most urgent challenges in the field. Digital microfluidic systems now allow for screening with as few as 3,000 cells per condition, making it feasible to work with limited primary animal cells that are not commercially available. Meanwhile, focused libraries like MyoCRISPR-KOLib efficiently target 90% of relevant transcripts while covering only a third of the genome [3][6]. This level of precision and efficiency is critical for overcoming resource constraints and scaling production.
"These findings demonstrate the utility of CRISPR screening for optimising bovine stem cell traits and offer a path toward more scalable cultivated meat production in the future." [1]
Despite these advancements, success hinges on access to the right infrastructure. Researchers need species-specific gRNA libraries, growth media designed for food applications, compatible bioreactors, and analytical tools tailored to cultivated meat production rather than pharmaceutical use. Addressing these needs, Cellbase has emerged as a vital resource, providing a transparent procurement system and a network of food-validated suppliers. By bridging gaps in resource accessibility, it helps streamline the path from CRISPR screening to industrial-scale production.
For teams working to engineer the next wave of cultivated meat cell lines, the tools and technologies are ready. The challenge now lies in the swift and effective deployment of CRISPR screening to realise its full potential.
FAQs
How do you choose between CRISPR knockout, CRISPRi and CRISPRa for a screen?
The choice between these systems hinges on your specific biological question and the outcome you're aiming for:
- CRISPR knockout: This method disrupts gene function entirely, making it ideal for studying the effects of gene loss or inactivation.
- CRISPRi: By repressing gene expression without cutting the DNA, this approach is well-suited for investigating essential genes or when reversible suppression is required.
- CRISPRa: If you need to upregulate gene expression, this system is the go-to choice. It's particularly useful for examining the effects of overexpression, such as promoting cell proliferation or differentiation.
When deciding, take into account your cellular model, the genes you’re targeting, and the overall goals of your experiment.
How can you boost proliferation without harming muscle or fat differentiation?
Boosting the proliferation of muscle or fat cells while maintaining their ability to differentiate is a key challenge in cultivated meat production. One promising approach involves CRISPR-based gene editing, which allows precise manipulation of genes to enhance growth or extend cell lifespan. For instance, targeting myostatin (MSTN) can promote cell growth, while editing CDKN2A helps cells bypass senescence.
That said, achieving a balance between proliferation and differentiation is critical. Mismanagement of certain targets, such as P53 (TP53), could impair differentiation, potentially compromising tissue quality. To navigate these complexities, high-throughput CRISPR screening is instrumental. This technique identifies the most effective gene regulators, paving the way for scalable and healthy tissue development in cultivated meat production.
What’s needed to validate CRISPR screen hits before scaling a cell line?
Validating CRISPR screen hits for cultivated meat production requires a methodical approach. First, gene function must be confirmed through independent experiments, such as gene knockouts, to ensure the observed effects are reproducible. Next, it's crucial to evaluate the biological relevance of these genes by examining their impact on factors like cell proliferation, viability, and longevity.
Safety assessments are equally important to rule out off-target effects or genetic instability that could compromise the process. Functional validation under conditions that mimic industrial settings, such as bioreactors, is another critical step. This ensures that the genetic edits perform as expected in large-scale production environments. Thorough testing at every stage is non-negotiable before considering scale-up.











